Separation of Nucleotides
In this application, we present an easy-to-use protocol for sensitive detection and quantification of nucleotides in red blood cell extracts. Retention times were very reproducible with %RSD around 0.4. The desired separation between UDP-Glu and UDP-Gal was achieved, all other nucleotides eluted later and did not interfere with the analysis. ATP represents the most abundant nucleotide in the red blood cell extracts and in this example it eluted after both analytes.
Figure A: Sample (spiked with UDP-Galactose) of a red blood cell extract from a patient for monitoring, Figure B: Standards using the same method.
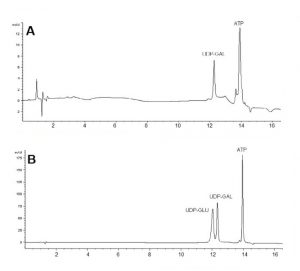
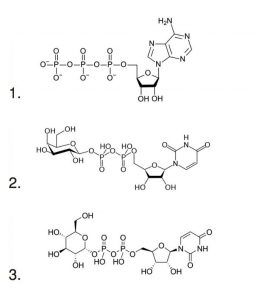 Peaks:
Peaks:
1. UDP GLU: UDP-Glucose: Uridine 5’-Diphosphateglucose, 1 mg/mL
2. UDP-GAL: UDP-Galactose: Uridine 5’-Diphosphategalactose, 1 mg/mL
3. ATP: Adenosine 5’-Triphosphate, 1 mM
Method Conditions
Column: Cogent Diamond Hydride™, 4μm, 100Å
Catalog No.: 70000-15P-2
Dimensions: 2.1mm x 150 mm
Mobile Phase:
—A: DI Water / 13mM Ammonium Acetate
—B: 90% Acetonitrile / 10% DI Water / 13mM Ammonium Acetate
| Time (minutes) | %B |
| 0 | 100 |
| 1 | 95 |
| 8 | 95 |
| 9 | 80 |
| 10.5 | 80 |
| 14 | 50 |
| 15 | 100 |
Post Time: 5 minutes
Injection vol.: 1 μL
Flow rate: 0.4 mL / minute
Detection: UV @ 254nm
Notes: Even with UV detection it is possible to detect individual nucleotides in the presence of a biological matrix. The retention time can be compared to the standard (Figure B) showing that the values of the real sample and the standard are very close. UDP-glucose, UDP-galactose and galactose 1-phosphate determination can be used for diagnosis of galactosemia in newborn babies [1-2].
1. Ji-Seon Jeong, Hye-Ran Yoon, Seon-Pyo Hong, Development of a new diagnostic method for galactosemia by high-performance anion-exchange chromatography with pulsed amperometric detection, J. Chromatography A, 1140 (2007) 157-162. 2. M.T. Matyska, J.J. Pesek, J. Duley, M. Zamzami, S.M. Fischer, Aqueous NormalPhase retention of nucleotides on silica hydride-based column: Method development strategies for analytes relevant in clinical analysis, J. Sep. Sci. 33 (2010) 930-938
Attachment
ATP, UDP-Glucose UDP-Galactose Separation from Red Blood Cells- pdf 0.2 Mb Download File


